Research subjects
Various technologies will be developed to omit or simplify the crystallization process, which is the bottleneck in X-ray crystal structure analysis.

Research subjects
Various technologies will be developed to omit or simplify the crystallization process, which is the bottleneck in X-ray crystal structure analysis.

Research contents
Based on the technology developed in the research subjects, new research methods and approaches will be proposed and demonstrated in drug discovery, foods, fragrances, and chemical research.

Education
Through research, we will educate students and foster young researchers.
Enzyme for “secondary” metabolism in a giant virus
A giant virus, as its name suggests, is a very large virus. It was common knowledge that a virus was so small that it could not be observed with an optical microscope previously. However, the discovery of a giant virus changed the concept of the virus. The size of the giant virus can reach more than 10 times that of conventional viruses (less than 0.1 μm), and the giant virus can be easily observed with an optical microscope.
The giant virus is not just huge. It has many interesting properties. One of them is that the giant virus has enzymes for metabolism. Metabolism normally happens in living organisms, and many viruses do not have the enzymes for metabolism.
Metabolism can be classified into two groups, primary metabolism and secondary metabolism. Primary metabolism is essential for the survival of living organisms, whereas secondary metabolism is not absolutely necessary. Primary metabolism of giant viruses has already attracted much attention. However, secondary metabolism of the giant viruses has not been investigated yet.
In this study, we analyzed an enzyme for the secondary metabolism in the giant virus. Secondary metabolism is very important. For example, most of useful compounds isolated as drug candidates from nature are produced by secondary metabolism.
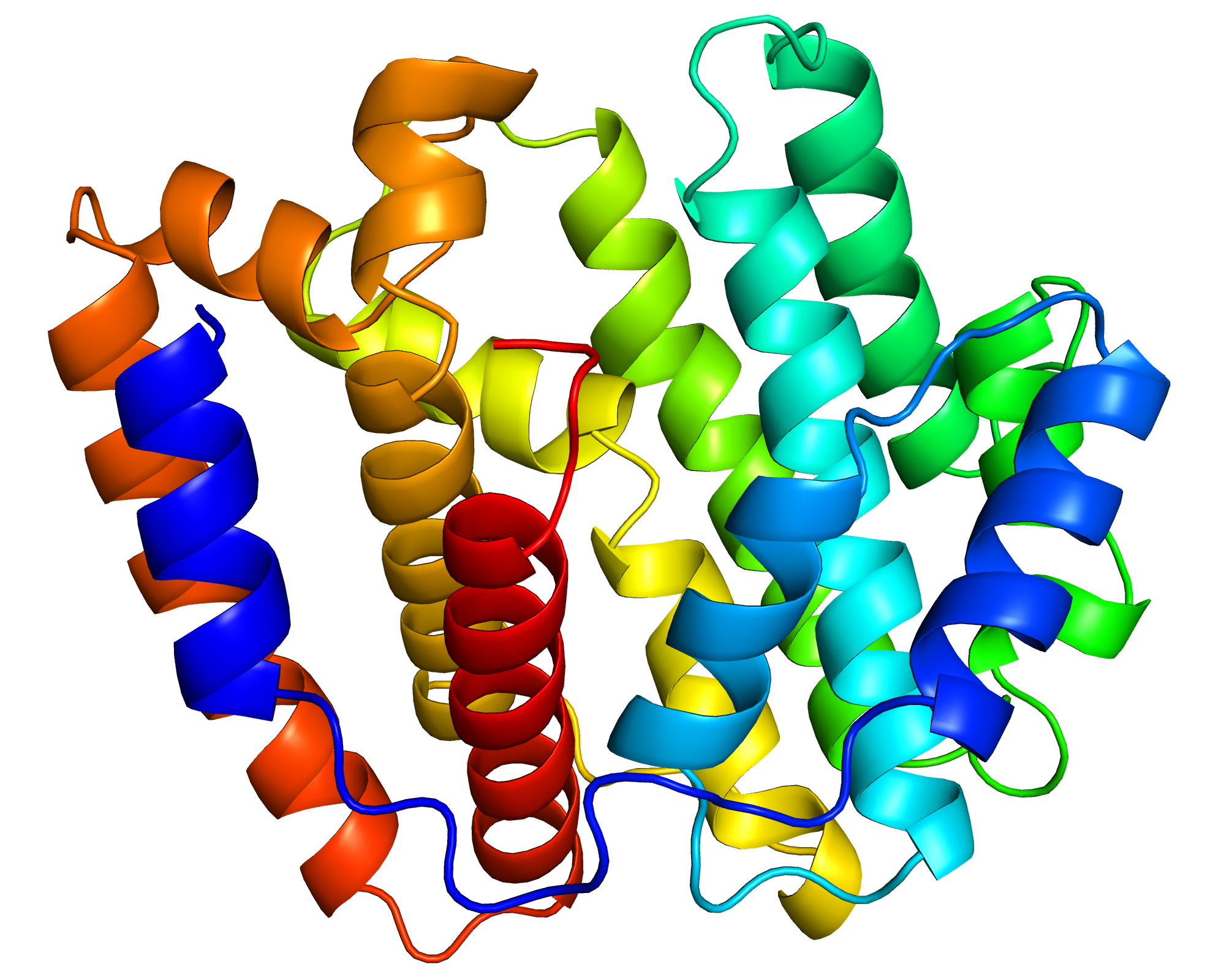
We focused especially on a terpene synthase, one of the important enzymes for secondary metabolism. We analyzed the function of this enzyme in detail. Moreover, we revealed the structure of this enzyme through the experiments at synchrotron radiation facilities. (The structure of the enzyme is shown in the figure above.) We believe that this study marks the beginning of studies on secondary metabolism in the giant viruses.
Function and Structure of a Terpene Synthase Encoded in a Giant Virus Genom
Y. Jung, T. Mitsuhashi, S. Sato, M. Senda, T. Senda, and M. Fujita
J. Am. Chem. Soc. 2023, 145, 25966-25970.
Biosynthesis of plant’s medicinal compounds: unveiling a 60-year-old mystery!
――The successful crystal structural analysis of unstable intermediates by the fusion of synthetic chemistry and supramolecular chemistry ――
[Abstract]
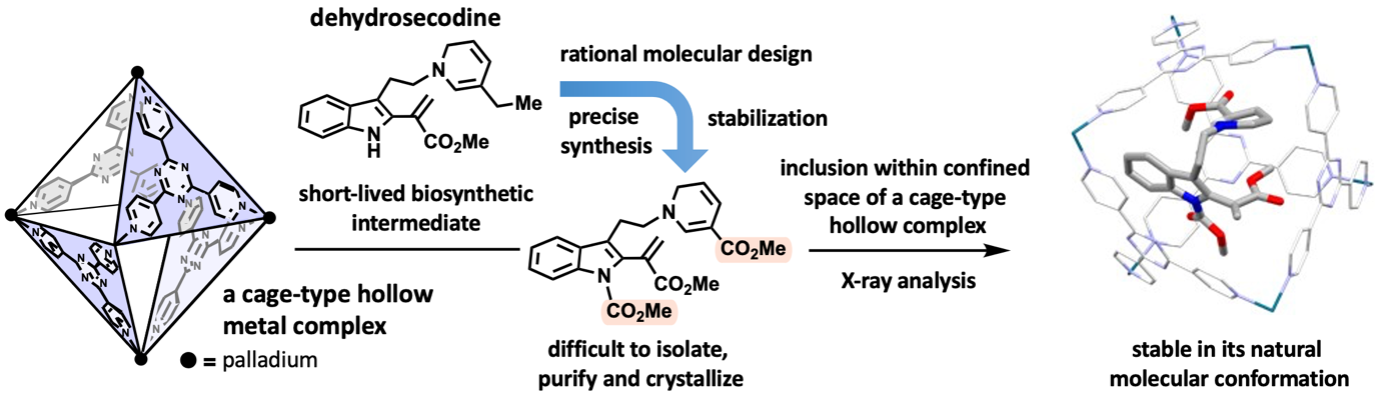
By combining natural products chemistry (Professor Hiroki Oguri, Graduate School of Science, The University of Tokyo) and supramolecular chemistry (Distinguished Professor Makoto Fujita, Project Professor Sota Sato, Dr. Hiroki Takezawa, Graduate School of Engineering, The University of Tokyo), a collaboration research work was reported in a scientific journal. The research team designed and synthesized stabilized molecules mimicking biosynthetic intermediates at the branching points of metabolic pathways in medicinally important plants and encapsulated them within hollow metal complexes. X-ray crystallography successfully clarified the three-dimensional structure. Dehydrosecodine, a hypothetical intermediate in the biosynthesis of vinca alkaloid anticancer drugs, was proposed more than 60 years ago. However, its three-dimensional structure had been a mystery due to its highly labile and short-lived nature. Oguri and co-workers designed and precisely synthesized a structurally modified intermediate by optimizing the reactivity and stability of dehydrosecodine. However, the stabilized intermediate remained difficult to isolate, purify, and crystallize. By exploiting a cage-type hollow complex developed by Fujita and co-workers. The labile intermediate was demonstrated to be stably trapped inside an isolated space in its natural conformation. X-ray crystallographic analysis of the inclusion complex revealed the molecule structure of dehydrosecodine-type compounds. This work is expected to be applied to the study of other biosynthetic processes via short-lived intermediates, and the results provides new insights into molecular transformations that generates skeletally diverse molecules.
[Detailed description]
Medicinally relevant plants biosynthesize a diverse range of monoterpene indole alkaloids. The iboga- and aspidosperma-type skeletons, which are building blocks of vinca alkaloid anticancer drugs, are biosynthesized from a common dehydrosecodine-type intermediate with versatile reactivities. Although the hypothetical dehydrosecodine intermediate existing at the branching point of the biosynthetic pathway has been proposed for approximately 60 years, its three-dimensional structure has remained a mystery due to it being extremely unstable and short-lived, and no one has isolated or determined its structure until now.
Oguri’s group designed and synthesized a moderately stabilized intermediate precisely mimicking dehydrosecodine. Although the stabilized dehydrosecodine-type intermediate was able to generate in flask, it was still too unstable to isolate, purify, and crystallize. In 2013. Fujita’s group invented the “crystalline sponge method” to determine the structure of small organic compounds in the internal pores of supramolecular metal complexes. This method has revolutionized the X-ray crystallography of organic compounds, which are generally difficult to crystallize.
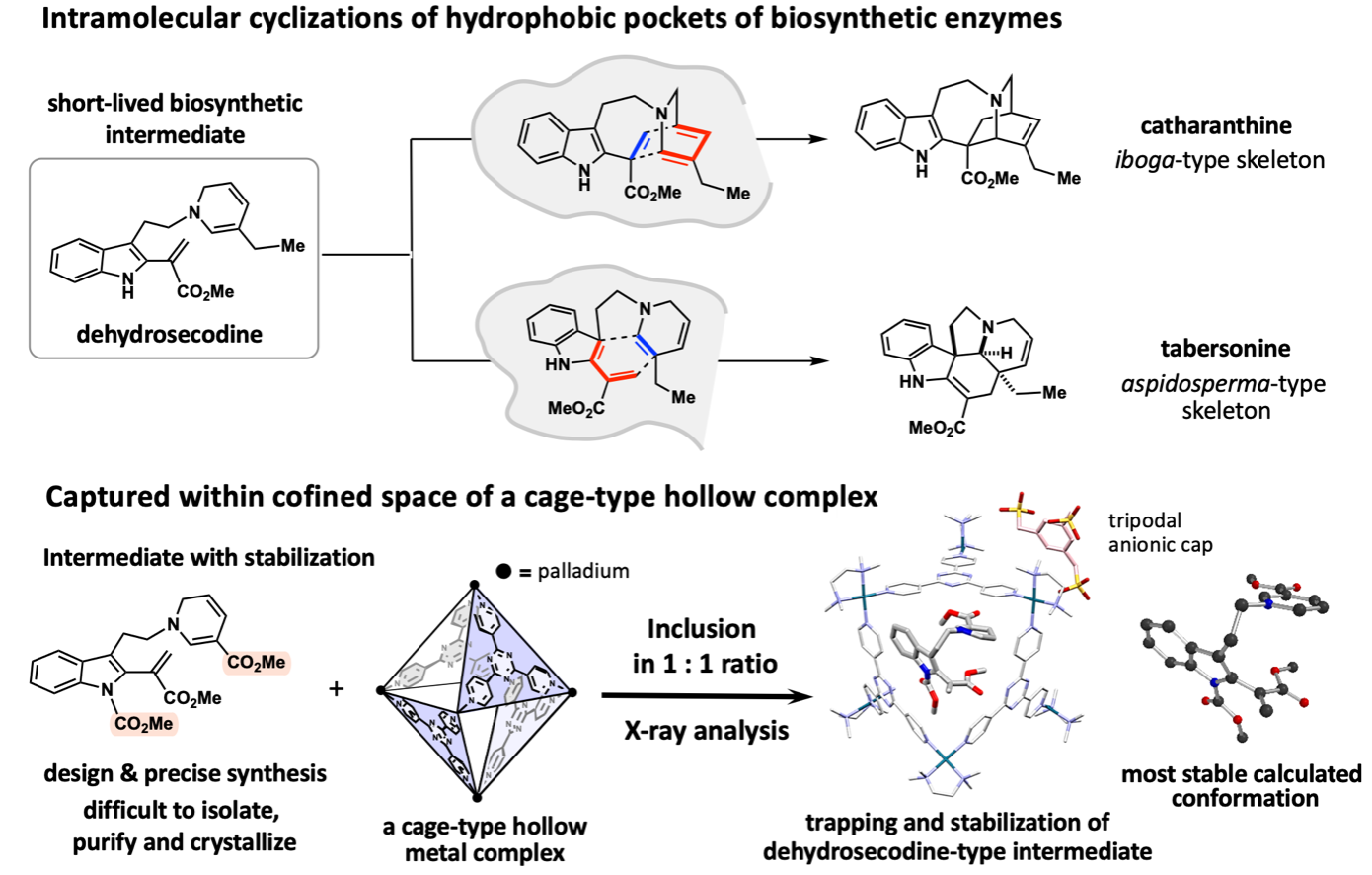
To achieve the X-ray crystallographic analysis by encapsulating and stabilizing a dehydrosecodine-type intermediate synthesized, the research team envisioned to make use of a hollow coordination complex developed in the Fujita group. Since the dehydrosecodine-type intermediate is relatively large in size, they utilized a cationic palladium complex with larger internal vacancy than the crystalline sponge. The unstable guest molecule entered the cage through the portal openings of the octahedral cage-shaped palladium complex. With investigation of the heating conditions and other factors for the efficient inclusion in a 1:1 molar ratio, the complexation in aqueous solution was confirmed by NMR analysis. The crystallization of the resulting complex was facilitated by coordinating the palladium cage with a tripodal anionic organic cap molecule, and crystals of the dehydrosecodine-type intermediate encapsulated within the cage framework were obtained. X-ray crystallography at SPring-8 BL26B1 beamline, a large synchrotron radiation facility, successfully clarified the three-dimensional structure of the dehydrosecodine-type intermediate, which has remained unresolved for many years.
Interestingly, the three-dimensional structure of the intermediate obtained in this X-ray crystallographic study is almost perfectly consistent with the most stable molecular conformation estimated by calculations. The dehydrosecodine-type compound was found to exist in a pre-folded state to undergo intramolecular cyclization reactions to construct the iboga- and aspidosperma-type skeletons. The results strongly support the involvement of dehydrosecodine as a common key biosynthetic intermediate for the divergent biosynthesis to various terpene indole alkaloids, a proposal made 60 years ago.
It was also found that the unstable and difficult-to-handle dehydrosecodine-type intermediates can be stored for a prolonged period of time when trapped within the cage complex. By trapping short-lived intermediates generated in the hydrophobic pocket of biosynthetic enzymes inside the isolated space of an artificial host, it provides an advanced research platform that enables the stable handling of short-lived molecules while observing the three-dimensional structure of unstable intermediates in its natural state.
This approach, which integrates natural products synthetic chemistry and supramolecular chemistry in a progressive manner, is expected to facilitate investigations of biosynthetic processes via short-lived intermediates. In addition, the synthetic approach of trapping dehydrosecodine-type intermediates, having a wide range of reactivities, inside a hollow host framework while stabilizing them, could provide deeper understanding of biosynthetic conversions and rational strategy for devising biomimetic transformations that generate skeletally diverse scaffolds reminiscent of natural products.
Synthetic Modulation of an Unstable Dehydrosecodine-type Intermediate and Its Encapsulation into a Confined Cavity Enable Its X-ray Crystallographic Observation
G. Tay, T. Wayama, H. Takezawa, S. Yoshida, S. Sato, M. Fujita, and H. Oguri
Angew. Chem. Int. Ed. 2023, in press.
The crystalline sponge method makes beer delicious!?

Beer is a drink that is loved all over the world and its refreshing bitterness is appreciated by many. Surprisingly however, the bitterness components contained in beer have not yet been completely elucidated scientifically. The bitter taste of beer originates from α-acid contained in the ingredient hop. During the brewing process, α-acid is isomerized to iso-α-acid, and it wasn’t until 2013 that the correct structures of these compounds were elucidated. This example illustrates the extreme difficulty of natural product structural analysis. It is known that α-acid and iso-α-acid are further chemically transformed into various compounds, but there are very few examples solving those structures. Thus there are still many compounds contained in beer that have not been chemically revealed. For the first time in the world we have succeeded in comprehensively elucidating the structure of trace amounts of unknown bitterness compounds by effectively combining HPLC and the crystalline sponge method. The components that were formed from iso-α-acid were found to possess complicated three-dimensional structures, which is where the crystalline sponge method demonstrated its effectiveness. By elucidating the structure of these reaction products, it becomes possible to understand the mechanism of the chemical change occurring. These findings enable control over the composition of bitterness components making it possible to control the reaction to adjust the taste and maintain freshness, leading to the development of new brewing technologies. This is an important achievement, we believe, in terms of the industrial applicability of the crystalline sponge method.
Comprehensive Structural Analysis of the Bitter Components in Beer by the HPLC-Assisted Crystalline Sponge Method
Y. Taniguchi, T. Kikuchi, S. Sato, and M. Fujita
Chem. Eur. J. 2021, 28, e202103339.
The extended sponge method – Aiming for protein structural determination through encapsulation in a gigantic cage
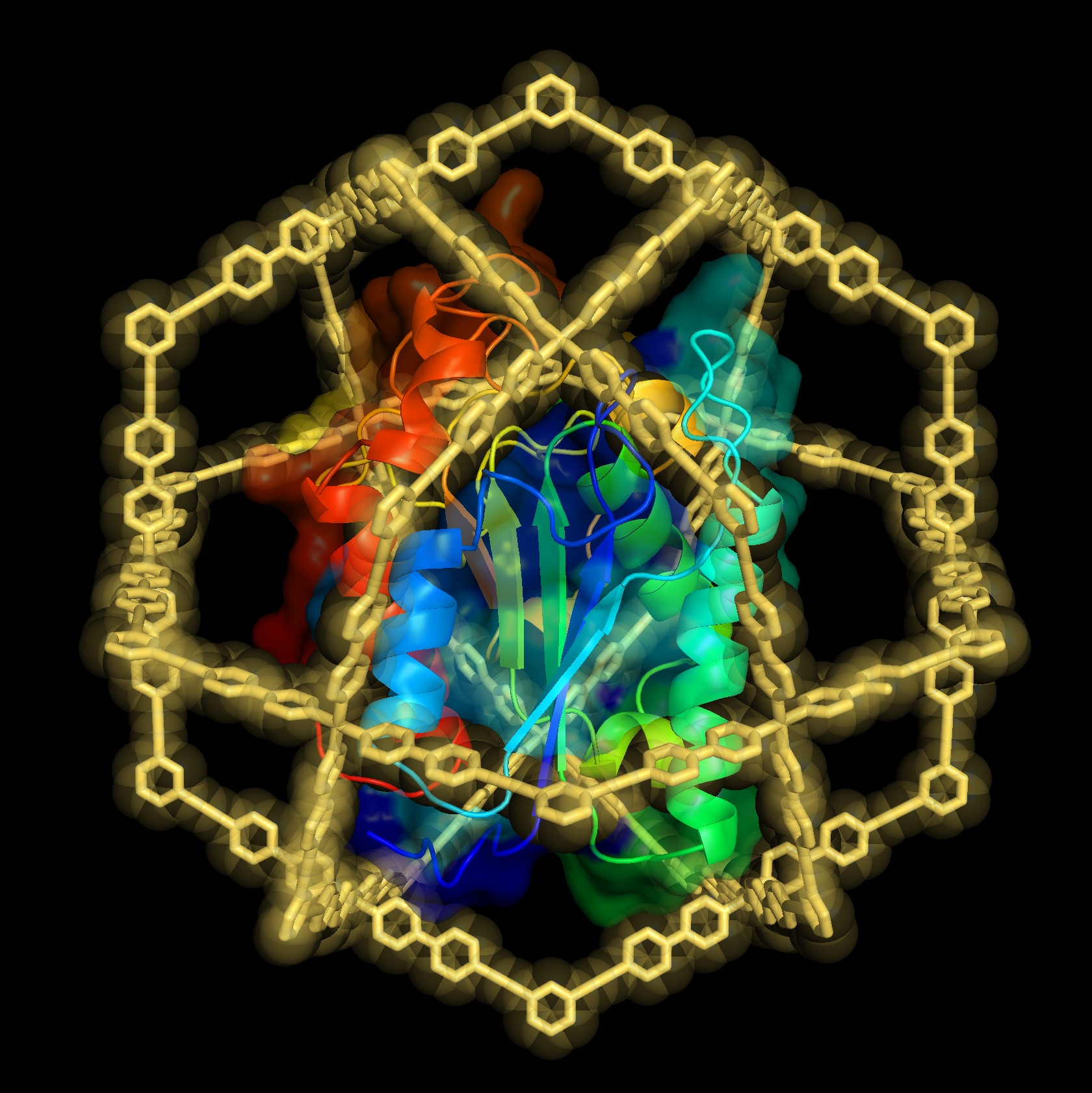

In this article, we used a self-assembled cage complex with a giant hollow inner space to precisely encapsulate a natural protein and analyzed its structure and properties in detail for the first time. We discovered that the protein is stabilized by confinement in its inner space and that refolding of the caged proteins can be induced, repairing the three-dimensional structure from a partially denatured state. By utilizing these “encapsulation effects” we will develop a method to elucidate protein structures difficult to observe by the conventional techniques.
“Does the behavior of proteins change by confining them into a narrow space?” To answer this question that has been raised out of curiosity in the study of self-assembly, we have spent the last 10 years studying the encapsulation of proteins in the hollow inner space of artificial molecules. Proteins (molecular weight >10,000) are significantly larger than small organic molecules (molecular weight <500), making it extremely difficult to encapsulate them in the hollow complex while maintaining their three-dimensional structure. Although we succeeded in constructing the world’s largest complex molecule with an inner space large enough to encapsulate a protein, what we could do at first was to just confirm the encapsulation in the complex by adding modification to a part of the protein (Nat. Commun. 2012, 3, 1093.).
In our research published in Chem, we achieved the encapsulation of a natural protein into the hollow complex without protein engineering by developing a new method. Through the selective condensation with the protein’s N-terminus, we encapsulated a protein retaining its native structure. What is more, we evaluated the structure and enzymatic activity of the encapsulated protein and elucidated the behavior of a protein confined in a narrow space for the first time. Since in our study we use the N-terminus, which is common to proteins, this research can be applied to almost all the natural proteins.
When we encapsulated a protein called cutinase-like enzyme (CLE) and analyzed its structure and function, an astounding stabilization of the protein was observed. For instance, in acetonitrile CLE is completely deactivated and loses its enzymatic function within one hour. In contrast, by encapsulation into the cage, the activity was retained even after one month (1000-fold stabilization). We found out that this is because in denaturing acetonitrile, the protein loses its higher-order structure, resulting in aggregation, whereas the encapsulated protein is spatially isolated, thus suppressing denaturation and aggregation. To our surprise, although the structure of the encapsulated CLE started to collapse after two weeks in acetonitrile, upon changing the solvent to water its higher-order three-dimensional structure was restored and its enzymatic activity regenerated. This refolding, which is reminiscent of molecular chaperones, occurred because we encapsulated just one protein molecule in the narrow space of the cage. In this way, it became clear that even for proteins “encapsulation effect” can take place, which triggers unique properties of proteins in a narrow space.
We call this protein encapsulation chemistry the “extended sponge method” and are continuing our research with the aim of developing it into a protein structure analysis technique. The defined inner cavity of the hollow complex provides a suitable place for obtaining proteins’ detailed structural information. What is more, we believe that by utilizing the “encapsulation effect,” it is possible to capture protein structures that no one was able to observe before. For instance, by capturing unstable proteins or intermediate states inside the hollow complexes, we can elucidate dynamic and transient protein structures that were difficult to observe until now. In the future, we envision developing a technique, a completely new approach to protein structural analysis that will innovate life science.
Protein stabilization and refolding in a gigantic self-assembled cage
D. Fujita, R. Suzuki, Y. Fujii, M. Yamada, T. Nakama, A. Matsugami, F. Hayashi, J.-K. Weng, M. Yagi-Utsumi, and M. Fujita
Chem 2021, 7, 2672–2683.
Crystalline Sponge method: Principle and proof-of-concept studies
This review describes the historical background and principles behind the birth of the crystalline sponge method, which is our core technology. In addition, the review touches on the technical improvements found by us and researchers in Japan and overseas since the development of the crystalline sponge method, and examples of effective application research of the crystalline sponge method.
Crystalline Sponge Method: X-ray Structure Analysis of Small Molecules by Post-Orientation within Porous Crystals—Principle and Proof-of-Concept Studies
N. Zigon, V. Duplan, N. Wada, and M. Fujita
Angew. Chem. Int. Ed. 2021, 60, 25204-25222.